Joint Center for Research in Sustainable Chemistry (Centro Conjunto de Investigación en Química Sustentable, CCIQS). Toluca, Estado de México, México.
Calixarenes as molecular recognition agents
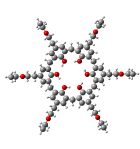
Calixarenes constitute a remarkable family of macrocyclic cavitands due to their functionalization possibilities and the array of sizes to which they can be synthesized. With a rather straightforward synthesis, calixarenes have drawn a lot of attention as molecular recognition agents. In our lab we calculate the different electronic interactions with various substrates so we can ultimately design a calixarene with tailor-made properties for molecular sensors, drug carriers and metal extraction agents.
Photosynthesis

Research on the Fenna-Matthews-Olsen complex in Green Sulfur Bacteria serves as a suitable model for the atomistic investigation about the first steps of energy transfer during Photosynthesis. The FMO acts as a molecular wire that transfers the energy from the pigment's excited states to the first reaction center in photosystem 2; recently, it has been discovered that the process by which this occurs is through coupled quantum coherence. Understanding of the photosynthesis process will allow us to eventually harvest sunlight more efficiently.
Non-canonical DNA

With the inclusion of d5SICS and dNaM into a modified strand of DNA in live E. coli bacteria, the realm of synthetic biology is faced with the challenge of getting new letters into the genetic alphabet that can lead to new ways we control gene expression. We use quantum mechanical calculations together with state-of-the-art molecular dynamics simulations to search the best candidates to join and interact with the ATCG group.
Sigma Holes in p-block atoms. Bonding properties & origins

Sigma holes are an anisotropy in the electrostatic potential of p-block atoms in which a region--opposite to a sigma bond--of positive potential, which can lead to typically negative atoms to behave as positive. This leads to the formation of halogen bonds and other kinds of secondary bonds (pnictogen, chalcogen, and aerogen bonds), but the nature of their electronic structure origins are still not clear.
Hello!, I am a chemistry teacher and PhD student in Turkey. I am now studing in synthesis chem (N-heterocyclic carbenes) But want to continue on computational chem. too. I don’t know which software to use and how to’s. So can you give a help to me about this. I want to learn Gaussian and how to make publishment from it. And i want to ask what about molekel… it is free and small but is it really practically suitable… Thank you.
Molekel is only a visualizer, that means you can’t perform calculations with it, only visualize the results from calculations performed elsewhere. I will post a short tutorial on Gaussian very soon (I promise) so you can learn how to start working with it, ok? Please stay tuned!! But let me tell you that learning all the things you can do with it can be a long process, so just be patient, ok? I have a friend here who works with N-heterocyclic carbenes, you could come to Mexico and work with her while you learn how to use Gaussian with me! how does that sound?
HAve a nice day!
” But let me tell you that learning all the things you can do with it can be a long process, so just be patient, ok?”
This has to be the understatement of the year! I am a junior Chemistry Undergrad and have been using Gaussian 03 & 09 for almost 16 months and I can wholeheartedly support the qoute as true. So, Good Luck!
Thanks Agapito!
At this rate, by the time you become a grad student you will be an expert on G09! Thank you for your comments here and on FB, perhaps we can meet one of these days, I will let you know next time I’m in Texas. In the mean time, take care and once again thanks for writing!
Professor, I think you should update your research group photo haha (just some random thoughts…:D, just kidding! :))
You are right, Dali! I totally should. But the thing is that Maru never lets me to get her picture, she is always making up excuses, like her hair not being ok and such, and Howard, well, Howard doesn’t come all that often haha but you are right, we should update that part of the blog. Thanks 🙂
haha but you are right, we should update that part of the blog. Thanks 🙂
Have a nice day!
Hi Professor. First I want to apologize cause i’ve could not take a look about all your publications in the blog. That’s why I ask you this: Can you give me more information over the researching in Photosyntesis you are carrying out now? Besides I want to know, how many posibilities are there for a posdoctoral scholarship in CCIQS. I’m chemist and a PhD student in physics (surface sciences); I’m working with VASP package making DFT calculations to describe chalcogens (S, Se) interactions with (111) silver’s face.
Hola Héctor!
The photosynthesis project is still in the developing stage, which is a fancy way to say we haven’t started it yet 🙂 Well, we have done some things but we are pretty much just setting the methodology now. This means there are open positions and opportunities for people interested in working with us!
Scholarships for postdocs can be found through the National University (which we belong to). I could get you more info if you are interested. We are interested in having valuable people working with us!
Have a nice day!
Hi Doc!
Have you received my email last week? just to check you are currently using that account.
Best Wishes!
Hello Hector!
I did get your email but Ive been terribly busy. I will send you some information about graduate studies here at CCIQS, ok?
Sorry for the delay!
Hi Dr. Thank you so much Your comments on solvation method were helped me more to run my Gaussian calculations. Best Wishes!
Glad I could help! 😉
Hi,
I am currently an undergraduate chemistry student studying in Scotland. I’m presently in my penultimate year and have some experience with computational chemistry (obtained by working in an Inorganic Computational Research Group last summer).
Would there be a possibility of an opportunity within your group this coming summer?
Regards,
Matthew.
Hello Matthew,
First of all thanks for being interested in our research group. About your question, yes you could come and work with us during the summer. We should organize our schedules since our university closes for a couple of weeks in July, but that can be arranged. The only problem is I don’t have a budget assigned for internships; I could arrange for some financing but I can’t promise anything at this point, therefore it would be much easier if you could find a source of financing from your own university. These are details we could take care of but the main answer is yes, we are interested in people working with us.
I look forward to hearing from you in the near future. Once again, thanks for your interest in our research
Best regards,
Thanks for the fast reply.
I will enquire within my department and get back to you asap!
Regards,
Matthew.
Hi,
Did you receive the email I sent you?
Regards,
Matthew.
Yes I did Matt, sorry if I haven’t replied; I will do so in a few minutes, plus I will try to find out if there is some source of financing I can find for you.
Best wishes
Hi Dr,
thanks in advance for your kindly help. my Q is how to use EMSL to get best initial gauss of basis set, for a system that it contains B and N atoms(almost 40 atoms ).
I’ve recently created a blog about Molecular Modelling. Researchers from computational chemistry background will find it pertinent. http://www.eventheodd.blogspot.in. Do follow and leave your comments.
It looks very nice, Chirag. Thanks for sharing it with us.
Have a nice day
Hai Sir,
I am using gaussian 09 for my phd work in nano clay modification studies.i want to use pbc.i am a beginner.i tried with examples from gaussian help for 1-D,2-D and 3-D.for 1-D &2-D it is working.but for 3-D it shows error.i tried some other 3-D examples also,but it is not running.we have only windows system.please give me some suggestions.
Thankyou Sir
Hello Thomas,
I need a little more information about the errors you’ve encountered so I can help you properly. Don’t hesitate to post your question again, please just do it more explicitly.
Have a nice day!
Hi Sir,
I try to calculate the interaction of Au(n=3,4) to methionine with MO6/SDD. I found that the interactions of Au-O and Au-OH can not optimized by l9999.exe. How can I sort out>
Thanks
Worasak
Dear Sir,
From Reference paper , I have read
A split valence (SV) of 433321/43321/43, they named the
SV4PPD basis set, where PP stands for two polarization functions
with exponents of 0.105 and 0.334.
How do give split valence (SV) of 433321/43321/43 for Iodine atom and the value of exponent in the input. And also how to decide the value of exponent for particular atom.
thanks in advance.
Ref:
Bull. Korean Chem. Soc. 2010, Vol. 31, No. 8 Chang Kon Kim et al.
DOI 10.5012/bkcs.2010.31.8.2228
Recently I was trying to calculate the effect of ligand field on the acetylenic C-H stretching frequency. But I am unable to understand how to calculate projection of electric field onto the corresponding bond generated by different ligands. I am using MP2 method under Gaussian09 for my calculation. It would have been very helpful if you guide me how to make input to specifically calculate the electric field potential at the coordinates of C and H atoms originated from ligand field.
Hello sir, I am a chemistry teacher. I am doing research in computational chemistry specifically on Irc calculation in gaussian 09. Could you please help me .
Yes of course. What is it that you need help with? There is a post on TS and IRC calculations in this blog. Try finding it and follow it.
I hope this helps
Dear Sir!
I am a chemistry teacher and PhD scholar in China. My major is computational chemistry my research project is on Computational insight for ring opening polymerization by N-heterocyclic carbenes (NHCs). Very recently I have changed my supervisor and switched to this field therefore I have not enough knowledge and understanding this field although I am trying my best.
Would you like to introduce any Professor related to my field? . If you are convenient I want to take your assistance in this regard as time to time. For future I am really motivated to create research collaboration with your group, if possible?
Best Regards.
Dear colleague
I am a computer scientist in the IPREM French chemistry lab located in Pau, France (http://iprem.univ-pau.fr/live/). One part of my job consists in installing on our cluster the softwares needed by the chemistry researchers of the lab.
Two versions of Molekel are used by my colleagues: the old 4.3 and the latest 5.4. They still need the 4.3 version for some treatments. Recently we went into some trouble when trying to make use of our old Linux 32-bit executable of Molekel 4.3 on our latest 64-bit architectures.
This is why I would like to try to compile Molekel 4.3 on a Linux 64-bit architecture. To do that, could you please provide us the source files of Molekel 4.3 ? I saw that you did this job for Win32 (https://joaquinbarroso.com/2012/01/11/molekel-4-3-win/).
I know that this version is outdated, but I had this demand from my collegues. We will use it at our risks, being aware that there is no support anymore for that version.
Thanks you in advance for your help.
Best regards.
Dear Marc,
I’m very sorry but I don’t seem to have those files anymore. Molekel’s use was deprecated by my team so long ago and we’ve changed equipments many times now that they seem to have been lost through the cracks.
I would encourage your colleagues to move to Chimera which is vastly superior to Molekel.
I hope this helps. Have a nice day!
Dear Joaquin
Thank you for your answer about Molekel. My colleagues like Molekel 4.3 because it allows to visualize very easily NBOs, which is not possible with the 5.4 version.
Do you know if it is possible to open .31~.47 files with Chimera ? Or with other softwares (maybe Chemcraft, Multiwfn…).
Thank you again for your help.
Best regards.
how to know from the energies of toluene and benzene that toluene is obtained from benzene?
I don’t think that’s possible
I would like known how to calculate ionisation potential and electronaffinity for monoanion system?
is this the same procedure as for the neutral system?
IP = E(cation) – E(neutral) and EA = (anion) – E(neutral)
Dear barroso,
I have run the job “# opt=calcall freq=raman rb3lyp/6-31g(d) nosymm guess=(mix,save) pop=nbo maxdisk=300GB geom=connectivity” in GaussianW 09 for many times but it outputs the below error message:
” Differentiating once with respect to nuclear coordinates.
There are 330 degrees of freedom in the 1st order CPHF. IDoFFX=5.
330 vectors produced by pass 0 Test12= 1.30D-13 1.00D-09 XBig12= 6.55D+02 7.17D+00.
AX will form 33 AO Fock derivatives at one time.
330 vectors produced by pass 1 Test12= 1.30D-13 1.00D-09 XBig12= 1.88D+02 2.09D+00.
328 vectors produced by pass 2 Test12= 1.30D-13 1.00D-09 XBig12= 1.16D+00 7.31D-02.
327 vectors produced by pass 3 Test12= 1.30D-13 1.00D-09 XBig12= 4.52D-03 2.81D-03.
326 vectors produced by pass 4 Test12= 1.30D-13 1.00D-09 XBig12= 6.47D-06 1.21D-04.
195 vectors produced by pass 5 Test12= 1.30D-13 1.00D-09 XBig12= 6.39D-09 3.82D-06.
3 vectors produced by pass 6 Test12= 1.30D-13 1.00D-09 XBig12= 4.99D-12 1.21D-07.
3 vectors produced by pass 7 Test12= 1.30D-13 1.00D-09 XBig12= 5.33D-15 3.20D-09.
Inverted reduced A of dimension 1842 with in-core refinement.
NIJ > Max2 in MMCore.
Error termination via Lnk1e in C:\G09W\l1002.exe at Thu Jun 13 01:55:34 2019.
Job cpu time: 0 days 14 hours 58 minutes 48.0 seconds.
File lengths (MBytes): RWF= 8921 Int= 0 D2E= 0 Chk= 86 Scr= 1″.
To over come this error message, what kind of Gauussian KEY(s) should I add to my input file ?
Dear Prof. Barroso,
I am trying to simulate a magnetic data in PHI software but I am facing problem. Please help me.
I am having a dimetallic bisporphyrin complex where one metalloporphyrin ring is connected with another metalloporphyrin ring through an ethane (-CH2-CH2-) bridge. Both the metal and porphyrin ring contains one unpaired electron in each. In one metalloporphyrin unit, the metal unpaired electron is antiferromagnetically coupled with the unpaired electron of the ring and the complex also shows some ferromagnetism.
Will I consider both antiferromagnetic and ferromagnetic coupling in the input file for magnetic simulation in PHI software??????
I was considering only antiferromagnetic coupling in the input file and tried to simulate it. But I failed to simulate it. Here I have attached my input file.
Can you please help me in making the input file???
Atoms numbering in the input file:
1 (one metal center), 3 (one ring center, coordinated with metal center 1)
2 (another metal center), 4 (another ring center, coordinated with metal center 2)
****spins
1
1
1
1
****fit
simplex
-1
ex 1 2 4
—-
-10
ex 1 3 4
ex 2 4 4
—-
-0.05
ex 3 4 4
—-
-40
cf 1 2 0
cf 2 2 0
—-
100
im 0 0 0
—-
0.008
ti 0 0 0
****params
opmode fit s
imp 1 1
****sus
bsus 0.100000
****end
hi dear Dr,
I am a drug delivery and drug discovery researcher and working on finding the intermolecular interactions between some quantomdots and bioagents using DFT methods. I am using G16 in ubuntu to run the calculations. I will appreciate it if you can help me evaluate the intermolecular interactions and compare them. which calculation should be run to find the interactions? and what factors show the strength or weakness of the interactions?
Hello sir,
I am working on covalent organic frameworks (COF). I want to computationally calculate binding energy of COF towards different metal ions.We have to optimise the COF and COF with metal ion.
Our COF consists of C,H,N,B,O and the metal ion calcium.We have tried with DFT
PBE-D3M/6-311G .
B3LYP/6-311G.
It failed to converge the structure.
Error messages appeared:
cphf not converged
Aborting SCF (increased accuracy required)
Aborting SCF due to too many large energy changes.
can you suggest better way of optimising our structure.