Coding, Computational Chemistry, DFT, Fukui, Gaussian, NBO, NBO, White papers
Tag: Fukui
3 Posts
3 Posts
Coding, Computational Chemistry, DFT, Fukui, Gaussian, NBO, NBO, White papers
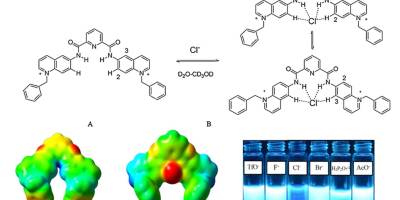
Chemosensors, Computational Chemistry, Paper, Sensors
Computational Chemistry, Theoretical Chemistry, White papers